Structure of AP-2 alpha appendage bound to the FxDxF motif of CCDC32
Sloan, D.E., Matthews, A.E., Tedamrongwanish, T., Nicely, N.I., Baker, R.W.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| AP-2 complex subunit alpha-2 | 244 | Mus musculus | Mutation(s): 0 Gene Names: Ap2a2, Adtab | 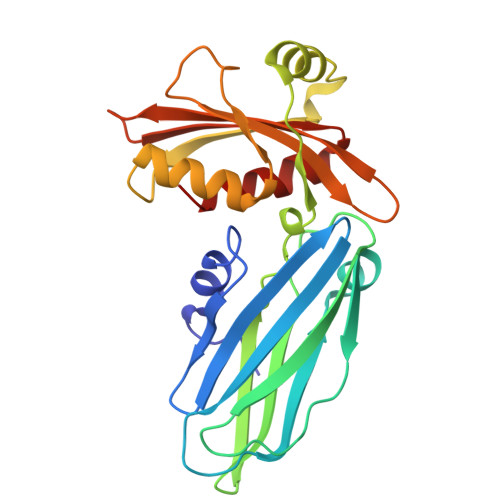 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P17427 (Mus musculus) Explore P17427 Go to UniProtKB: P17427 | |||||
IMPC: MGI:101920 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P17427 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Coiled-coil domain-containing protein 32 | B [auth P] | 9 | Mus musculus | Mutation(s): 0 | 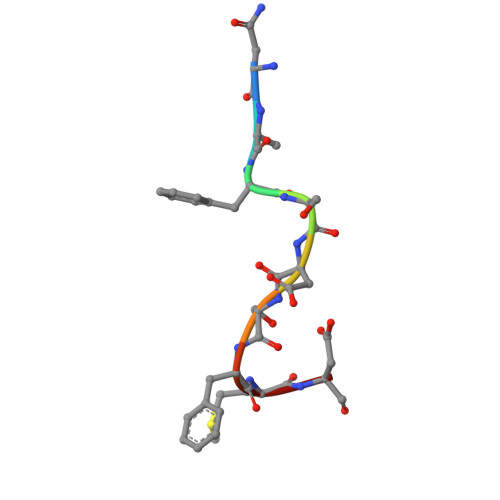 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q8BS39 (Mus musculus) Explore Q8BS39 Go to UniProtKB: Q8BS39 | |||||
IMPC: MGI:2685477 | |||||
Entity Groups | |||||
| UniProt Group | Q8BS39 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 40.807 | α = 90 |
| b = 72.465 | β = 99.092 |
| c = 42.053 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| SAINT | data reduction |
| SADABS | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | R35GM150960 |