The desensitized state structure of human P2X2 receptor channel in lipid nanodiscs with free ATP and sodium
Dhingra, S., Swartz, K.J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| P2X purinoceptor 2 | 471 | Homo sapiens | Mutation(s): 0 Gene Names: P2RX2, P2X2 | 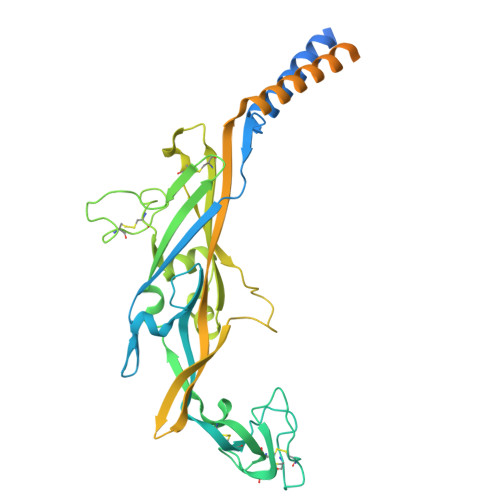 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q9UBL9 (Homo sapiens) Explore Q9UBL9 Go to UniProtKB: Q9UBL9 | |||||
PHAROS: Q9UBL9 GTEx: ENSG00000187848 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9UBL9 | ||||
Glycosylation | |||||
| Glycosylation Sites: 3 | Go to GlyGen: Q9UBL9-1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ATP (Subject of Investigation/LOI) Query on ATP | G [auth A], K [auth B], O [auth C] | ADENOSINE-5'-TRIPHOSPHATE C10 H16 N5 O13 P3 ZKHQWZAMYRWXGA-KQYNXXCUSA-N |  | ||
| NAG (Subject of Investigation/LOI) Query on NAG | D [auth A] E [auth A] F [auth A] H [auth B] I [auth B] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | 1.20.1_4487: |
| RECONSTRUCTION | cryoSPARC |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of Neurological Disorders and Stroke (NIH/NINDS) | United States | NS003018 |