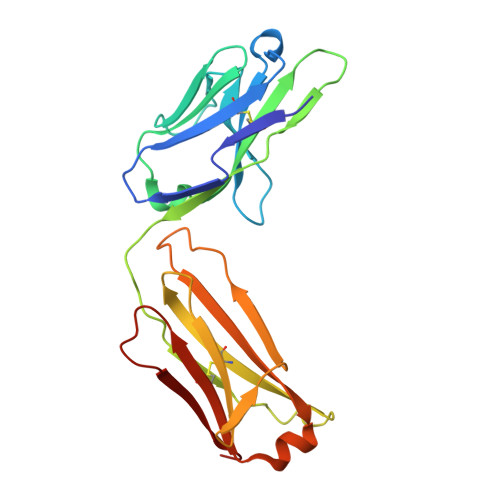
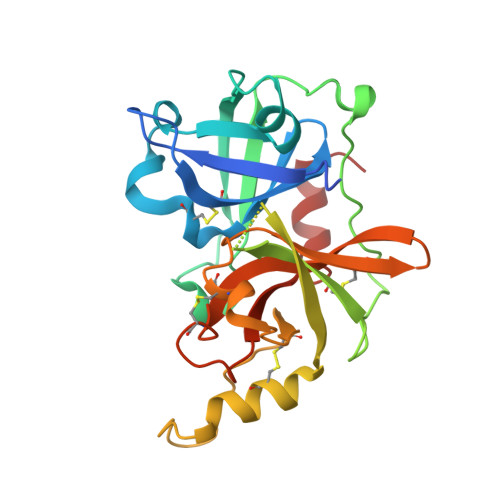
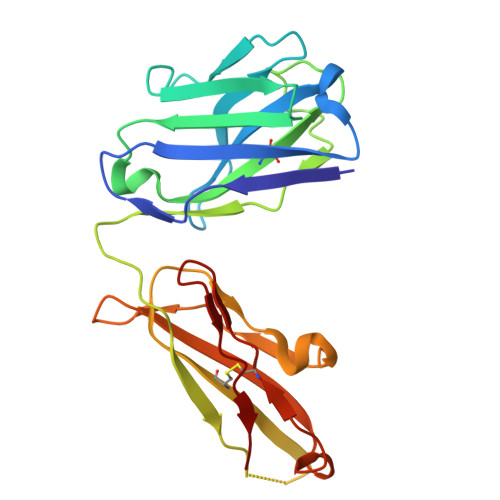
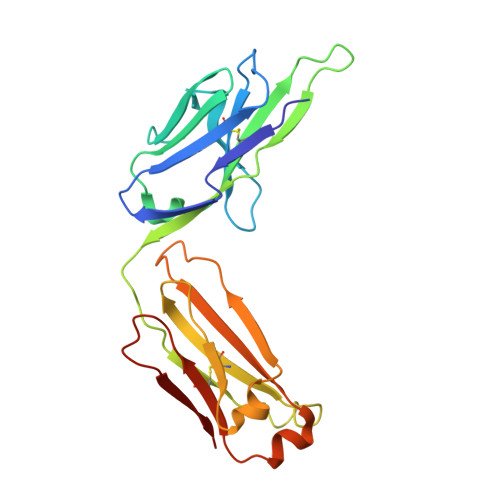
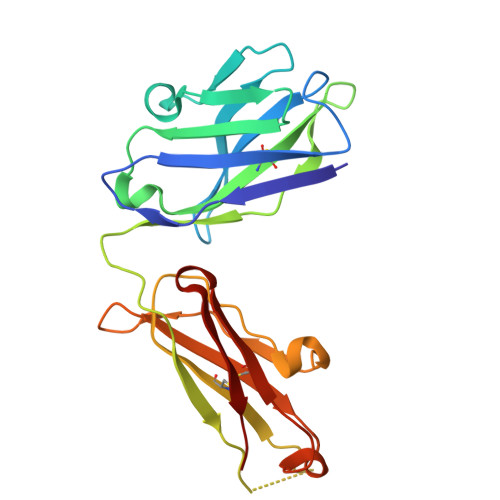
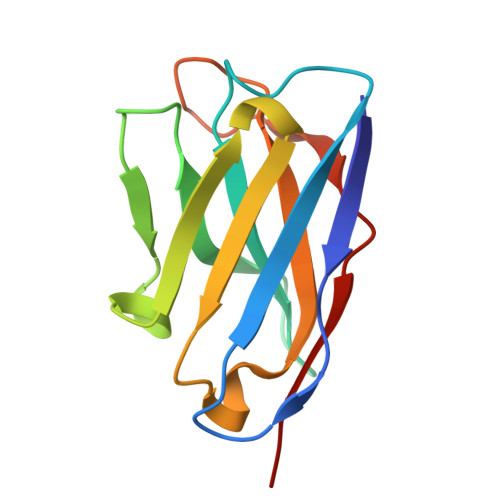
Complete inhibition of beta-tryptase by tetramer dissociation and active site allostery due to a single antibody residue.
Maun, H.R., Azumaya, C.M., Walters, B.T., Vij, R., Morando, A., Loyet, K.M., Koerber, J.T., Rohou, A., Lazarus, R.A.(2026) Nat Commun 17
- PubMed: 41957026
- DOI: https://doi.org/10.1038/s41467-026-70491-3
- Primary Citation Related Structures:
9MNB - PubMed Abstract:
Human β-tryptase, a tetrameric trypsin-like serine protease, is an important mediator of inflammatory responses in asthma, allergy and other diseases. Here we report an anti-β-tryptase antibody with a superior mechanism of action compared to others since it not only inhibits tetrameric β-tryptase, but also completely inhibits monomeric β-tryptase activity. The antibody binds to an exosite that causes tetramer dissociation as either an IgG or Fab and, in addition, allosterically alters the substrate binding cleft on monomers, thus preventing substrate binding and proteolysis. We solve the cryoEM structure of the complex, generate biochemical data and engineer point mutations to elucidate the allosteric path of inhibition. This ultimately reveals a single Asp to Gly mutation in CDR-L3 that only slightly impacts binding affinity, but completely eliminates inhibitory activity. Finally, we improve antibody inhibitory potency up to 4.7-fold by structure-based design creating new charge-charge interactions. This antibody may have enhanced efficacy and potential to assess the relevance of β-tryptase, including monomers, in biological and clinical settings.
- Department of Early Discovery Biochemistry, Genentech, Inc., South San Francisco, CA, USA. maun.henry@gene.com.
Organizational Affiliation: