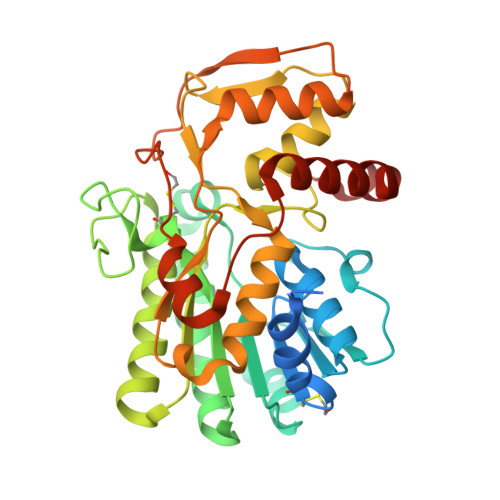
Structural characterisation of nucleotide sugar short-chain dehydrogenases/reductases from the thermophilic pseudomurein-containing methanogen Methanothermobacter thermautotrophicus Delta H.
Carbone, V., Schofield, L.R., Edwards, P.J.B., Sutherland-Smith, A.J., Ronimus, R.S.(2025) FEBS J
- PubMed: 40903920
- DOI: https://doi.org/10.1111/febs.70248
- Primary Citation of Related Structures:
6PMH, 6PNL, 8W3U, 9AR1 - PubMed Abstract:
Epimerases and dehydratases are widely studied members of the extended short-chain dehydrogenase/reductase (SDR) enzyme superfamily and are important in nucleotide sugar conversion and diversification, for example, the interconversion of uridine diphosphate (UDP)-linked glucose and galactose. Methanothermobacter thermautotrophicus contains a cluster of genes, the annotations of which indicate involvement in glycan biosynthesis such as that of cell walls or capsular polysaccharides. In particular, genes encoding UDP-glucose 4-epimerase related protein (Mth375), UDP-glucose 4-epimerase homologue (Mth380) and dTDP-glucose 4,6-dehydratase related protein (Mth373) may be involved in the biosynthesis of an unusual aminosugar in pseudomurein. In this paper, we present the structures of Mth375, an archaeal sugar epimerase/dehydratase protein (WbmF) determined to a resolution of 2.0 Å. The structure contains an N-terminal Rossmann-fold domain with bound nicotinamide adenine dinucleotide hydride (NADH) and a C-terminal catalytic domain with bound UDP. We also present the structure for Mth373 co-crystallised with uridine-5'-diphosphate-xylopyranose to a resolution of 1.96 Å as a NAD + -dependent oxidative decarboxylase (UDP-xylose synthase; EC4.1.1.35). Molecular modelling has also allowed for the identification of Mth380 as a UDP-N-acetylglucosamine 4-epimerase (WbpP; EC5.1.3.7), Mth631 as a UDP-glucose 4-epimerase (GalE; EC5.1.3.2) and Mth1789 as a classical dTDP-d-glucose 4,6-dehydratase (EC4.2.1.46). The UDP-sugar specificity of each archaeal nucleotide sugar short-chain dehydrogenase/reductase (NS-SDR) was elucidated via sequence, molecular modelling and structural analyses. Overall, these structures potentially shed light on the formation of the glycan portion of pseudomurein and capsular polysaccharide in Archaea.
- AgResearch Ltd., Grasslands, Palmerston North, New Zealand.
Organizational Affiliation: