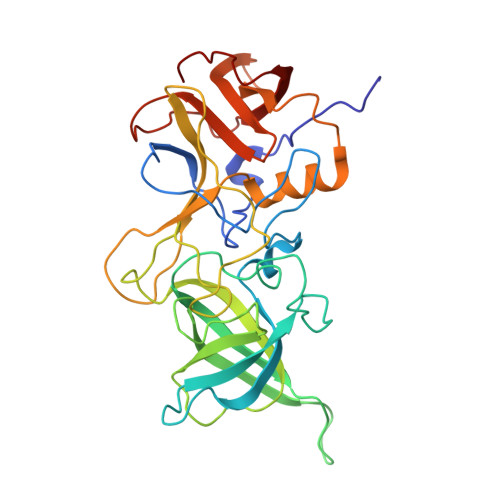
GII.23/24/25 noroviruses recognize glycans via a conventional glycan-binding site.
Li, H., Cong, X., Sun, X., Qi, J., Li, X., Jin, M., Duan, Z.(2026) Front Microbiol 17: 1767002-1767002
- PubMed: 41853721
- DOI: https://doi.org/10.3389/fmicb.2026.1767002
- Primary Citation Related Structures:
22WZ, 22XH, 22ZV, 22ZW - PubMed Abstract:
Human noroviruses (HuNoVs) are genetically diverse RNA viruses that cause acute gastroenteritis, with genogroup II (GII) accounting for over 90% of global infections. Glycans, particularly histo-blood group antigens (HBGAs), have been identified as attachment factors or receptors for HuNoVs infection. However, the glycan-binding receptors of the later-identified GII genotypes GII.23/24/25 remain elusive. We used saliva- and glycan-based ELISA assays to identify the binding spectra of GII.23/24/25 strains. We also solved the crystal structures of their P domains, including the GII.25 P domain in complex with the H disaccharide. Single-point mutagenesis was performed to identify key residues involved in glycan binding. The P domains of GII.24 and GII.25 can recognize multiple types of saliva samples, including both A/B/O secretor and nonsecretor individuals. In contrast, GII.23 primarily binds to B secretor saliva samples. Furthermore, GII.23/24 P domains are able to interact with the H disaccharide, whereas GII.25 exhibits binding affinity for both H disaccharide and B trisaccharide. Crystal structures of GII.23/24/25 P domains revealed high structural similarity, and the complex of GII.25 P domains with H disaccharide was resolved. Single-point mutagenesis identified N352, R353, D382, G443, G444, and H445 as critical residues for H disaccharide binding in GII.25 P domain, while A351 determines glycan-binding specificity. Our findings demonstrate that GII.23/24/25 exhibit glycan-binding patterns similar to most other GII HuNoV genotypes. The structural insights provide a better understanding of virus-host evolution and inform the development of therapeutic strategies against HuNoVs.
- Guangdong Provincial Key Laboratory of Agro-Animal Genomics and Molecular Breeding, College of Animal Science, South China Agricultural University, Guangzhou, China.
Organizational Affiliation: