Exploration of a workflow for the classification and identification of co-purified protein complexes yields new structures and multiple MSP assembly states.
Zhang, Q., Murthy, A.V., Mim, C.(2026) PLoS One 21: e0340207-e0340207
- PubMed: 41592050
- DOI: https://doi.org/10.1371/journal.pone.0340207
- Primary Citation Related Structures:
9HWJ, 9HWK, 9HWL - PubMed Abstract:
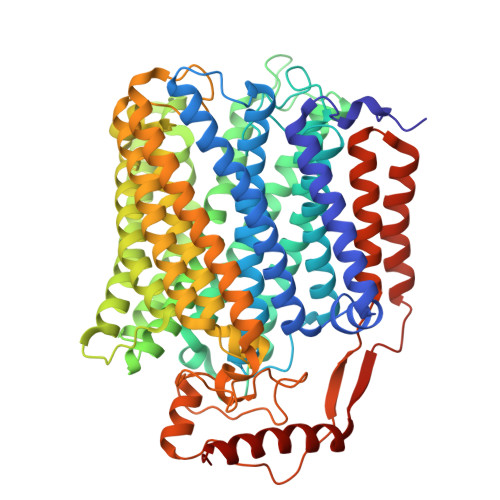
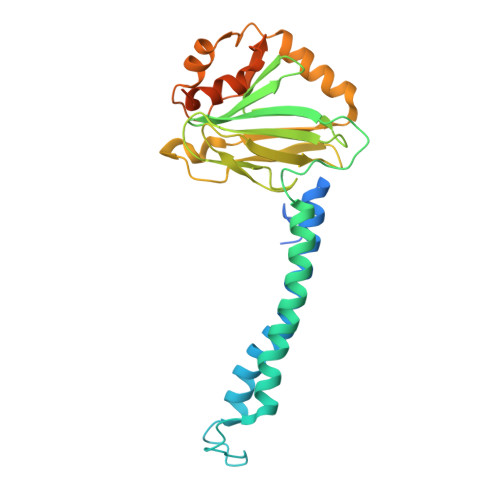
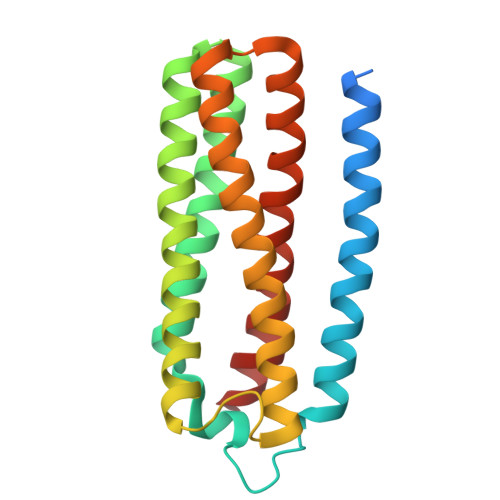
Native protein complexes have garnered interest as targets for structural dissemination. Cryogenic electron microscopy (cryo-EM) with its ability to image protein mixtures is the most promising tool to enable structural proteomics. Additionally, image processing has evolved and can deal with conformational and compositional heterogeneity. Integrative approaches, namely mass spectrometry in conjunction with cryo-EM, have made it possible to characterize and identify complex mixtures. However, this comes at a cost of generating models and interpreting mass spectra. Here we present a modified approach that builds on publicly available software. By generating maps around 4 Å and unsupervised model building we were able to identify the most abundant proteins in our sample. This sample consisted of co-purified membrane proteins in nanodiscs. We found a novel structure and unexpected nanodisc assemblies. Our maps imply a direct interaction between membrane proteins and membrane scaffolding proteins.
- Department of Protein Science, Kungliga Tekniska Högskolan (KTH)Stockholm, Sweden.
Organizational Affiliation: