Crystal Structure of Saccharomyces cerevisiae Sec14p in complex with phosphatidylcholine
Green, S.M., Laganowsky, A., Krieger, I., Singh, P.K., Sacchettini, J., Igumenova, T.I., Bankaitis, V.A.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| SEC14 cytosolic factor | 304 | Saccharomyces cerevisiae | Mutation(s): 0 Gene Names: SEC14, PIT1, YMR079W, YM9582.04 | 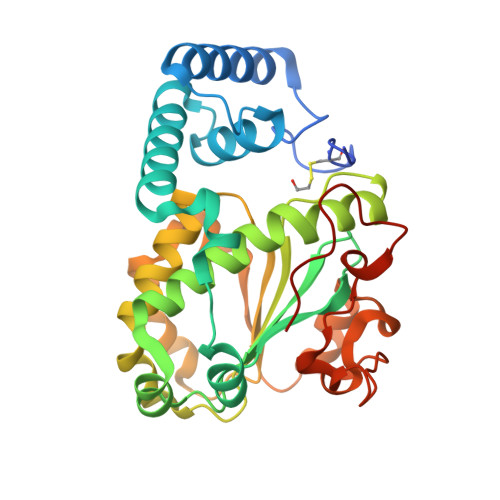 | |
UniProt | |||||
Find proteins for P24280 (Saccharomyces cerevisiae (strain ATCC 204508 / S288c)) Explore P24280 Go to UniProtKB: P24280 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P24280 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| POV (Subject of Investigation/LOI) Query on POV | C [auth A], F [auth B] | (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate C42 H82 N O8 P WTJKGGKOPKCXLL-PFDVCBLKSA-N |  | ||
| PEG (Subject of Investigation/LOI) Query on PEG | G [auth B], H [auth B], I [auth B] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| GOL (Subject of Investigation/LOI) Query on GOL | D [auth A], J [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| EDO (Subject of Investigation/LOI) Query on EDO | E [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| CME Query on CME | A, B | L-PEPTIDE LINKING | C5 H11 N O3 S2 |  | CYS |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 74.4 | α = 90 |
| b = 74.4 | β = 90 |
| c = 195.86 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |
| MOLREP | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Welch Foundation | United States | A-2194-20240404 |
| Robert A. Welch Foundation | United States | BE-0017 |
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | NIH R35 GM131804 |
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | NIH RO1 GM108998 |