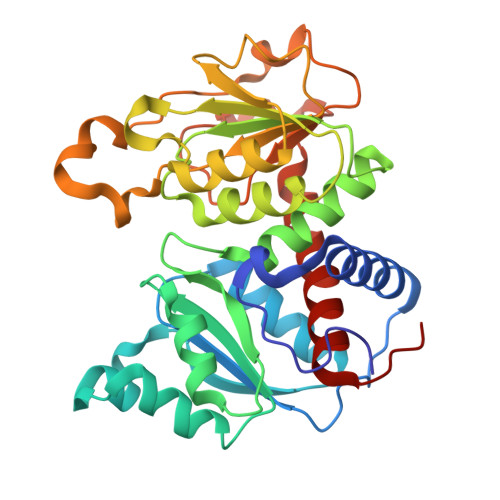
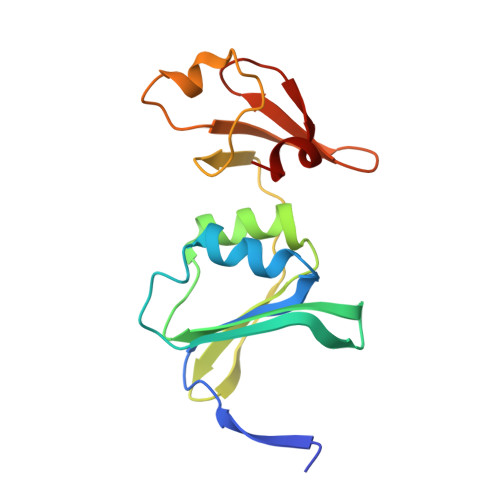
Cooperativity in E. coli aspartate transcarbamoylase is tuned by allosteric breathing.
Miller, R.C., Patterson, M.G., Bhatt, N., Pei, X., Ando, N.(2026) Nat Commun
- PubMed: 41862478 Search on PubMed
- DOI: https://doi.org/10.1038/s41467-026-70909-y
- Primary Citation Related Structures:
9EEH, 9EEJ, 9EEK, 9EEL, 9EEM, 9EEN, 9EEO, 9EEP, 9EEQ, 9EER, 9EES, 9EEU - PubMed Abstract:
Aspartate transcarbamoylase (ATCase) from Escherichia coli catalyzes a key step in pyrimidine nucleotide biosynthesis and has long served as a model for allosteric regulation. Despite decades of study, how nucleotide binding at distant regulatory sites controls cooperativity between active sites remained unresolved. Here we show that ATCase does not simply interconvert between two conformations, as traditionally depicted, but instead samples a continuum of conformations that tune enzyme cooperativity. Using complementary cryo-electron microscopy, small-angle X-ray scattering, and crystallography under conditions that ensure full assembly of the allosteric sites, we show that ATCase behaves like a flexible balloon whose global "breathing" motions directly regulate activity: compression enforces high cooperativity, inhibiting the enzyme, whereas expansion relieves this cooperativity and activates the enzyme. We further show that all four ribonucleoside triphosphates act in symmetric pairs to tune this motion, with the pyrimidines CTP and UTP compressing the enzyme to limit further pyrimidine production, and the purines ATP and GTP expanding it to balance pyrimidine and purine pools. Together, these findings uncover a dynamic breathing mechanism for long-range allosteric communication in ATCase.
- Department of Chemistry and Chemical Biology, Cornell University, Ithaca, NY, USA.
Organizational Affiliation: